16S Structured R Workflow and Complete 16S Pipeline for Microbiome Analysis
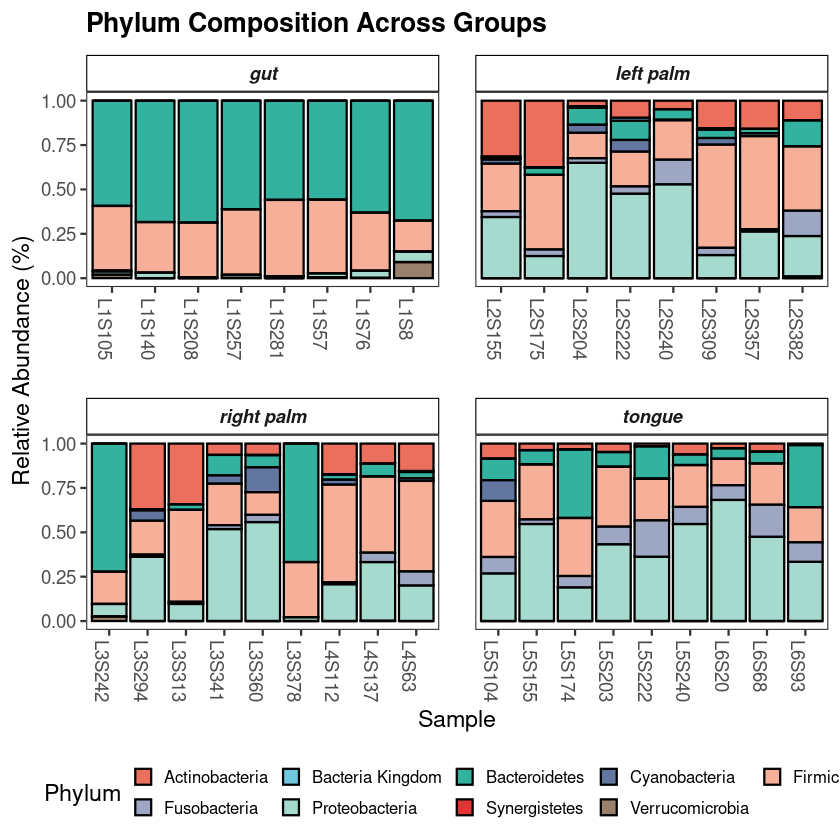
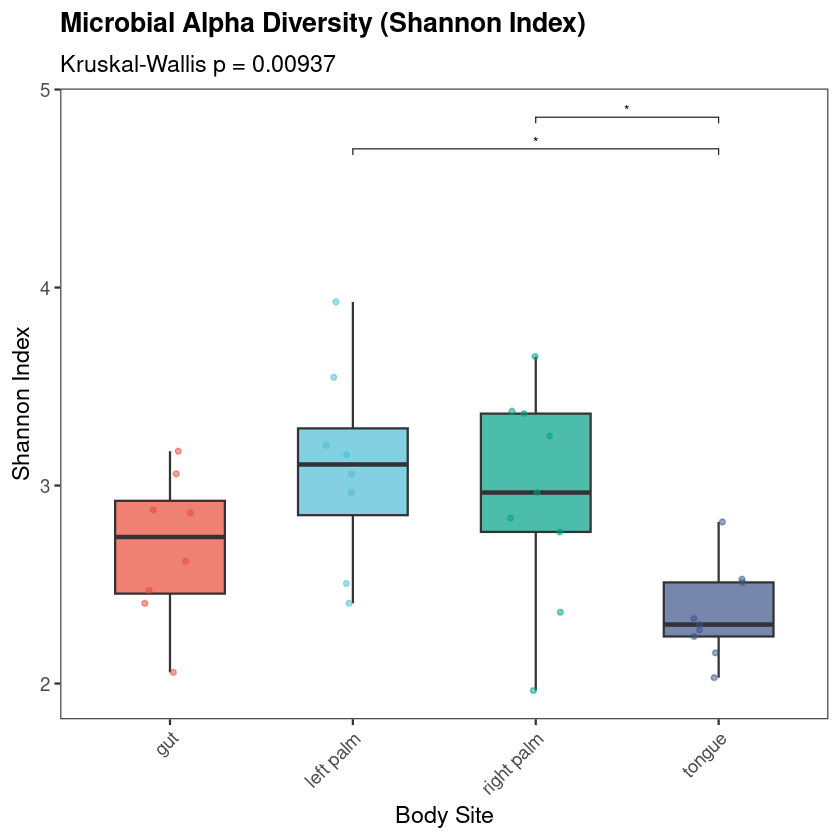
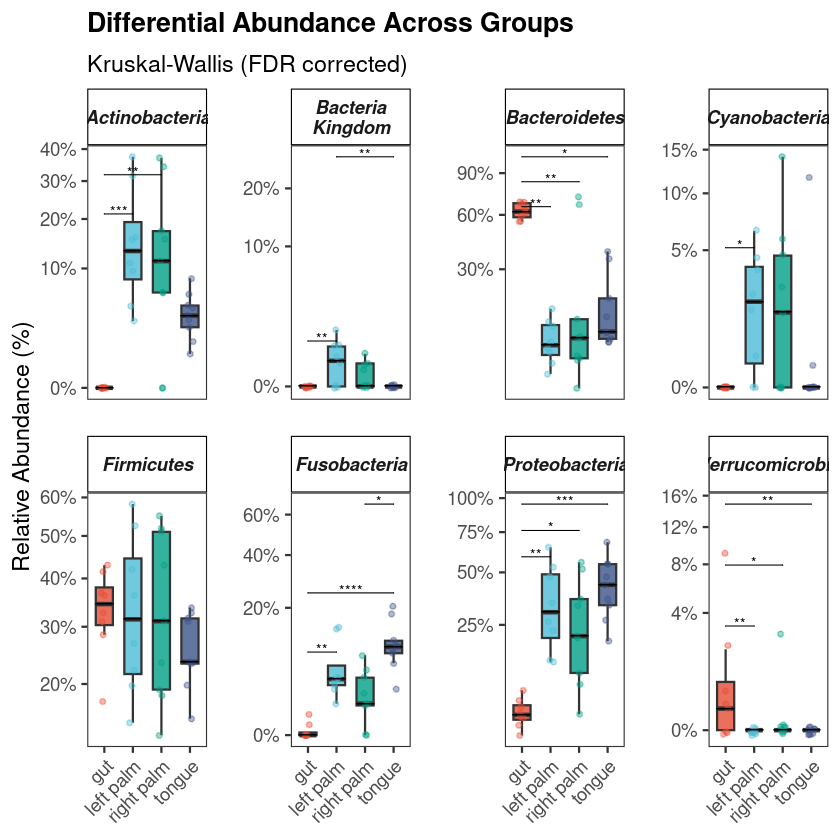
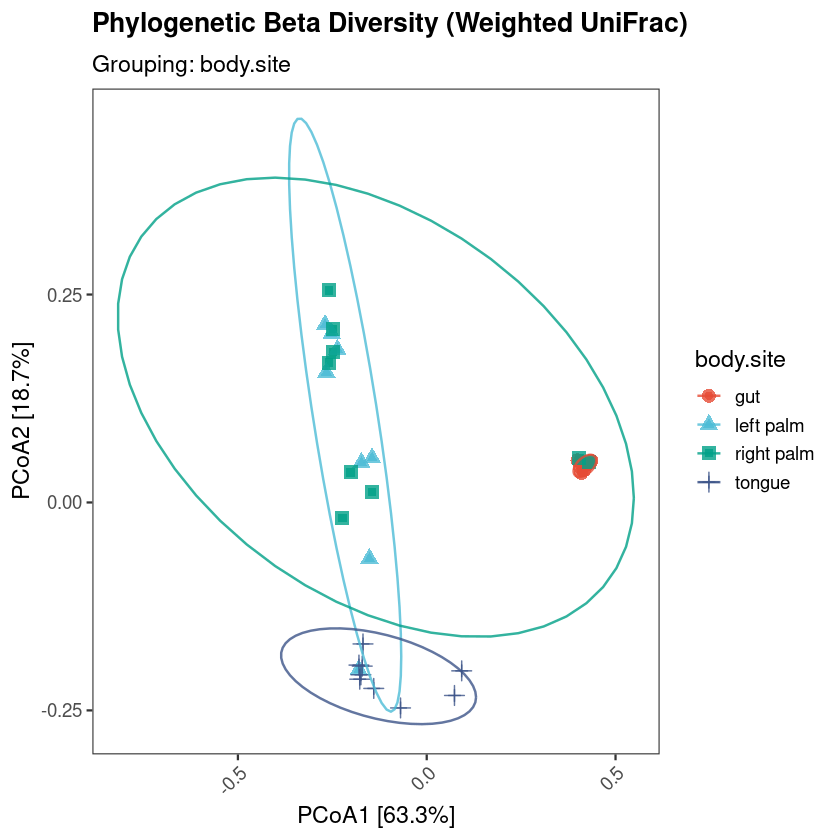
Get a ready-to-use 16S R code workflow and complete 16S pipeline for NGS meta-barcoding data downstream analysis and visualization in R. The script generates analysis-ready figures, exported CSV files, and a Methods section for microbiome projects.
Instant download. Individual license. No subscription.